AI Medical Record Parsing: CCDA XML to Health Data
How I built an AI medical record parsing system that extracts meds, labs, and allergies from CCDA XML and 30 MyChart PDFs into actionable health data.
By Mike Hodgen
My grandmother had seven specialists. Cardiologist, nephrologist, endocrinologist, pulmonologist, primary care, and two more I'm probably forgetting. Each one operated in their own silo. Each one had fragments of her medical history, but nobody — not a single provider — had the complete picture.
That's the problem I set out to solve when I built a health monitoring system for a family member with a similarly complex medical history. AI medical record parsing wasn't something I went looking for as a project. It was the only way to get a unified view of someone's health when the healthcare system refused to provide one.
The raw materials were all there. MyChart lets you download your records. Specialists will send PDFs if you ask nicely. But "access" and "usable" are two very different things. What I had was a stack of CCDA XML exports from Epic, 30+ PDFs from various providers, and a growing sense that no human being should have to manually reconcile all of this information.
This article is a technical build log. I'm going to walk through exactly how I parsed CCDA XML files, vectorized clinical PDFs, and built a RAG pipeline that can actually answer questions about a patient's medical history. If you're dealing with messy, high-stakes data in any industry, the pattern applies well beyond healthcare.
CCDA XML: The Healthcare Data Format Nobody Designed for Humans
What CCDA Actually Contains
CCDA — Consolidated Clinical Document Architecture — is the standard export format from electronic health record systems like Epic. When you download your records from MyChart, this is what you get.
In theory, it's structured. The XML contains discrete sections for medications, allergies, lab results, active problems, procedures, immunizations, vital signs, and more. Each section is identified by a templateId (an OID — object identifier) that tells you what kind of data lives inside.
In theory, this should be straightforward to parse. In practice, it's a nightmare.
Why It's Both Structured and Chaotic
A single medication entry in a CCDA file can span 40+ lines of nested XML. You'll find SNOMED codes, RxNorm codes, display names, dosage information, and status flags scattered across nested elements and attributes. The structure exists. The consistency doesn't.
 CCDA XML Inconsistency Problem
CCDA XML Inconsistency Problem
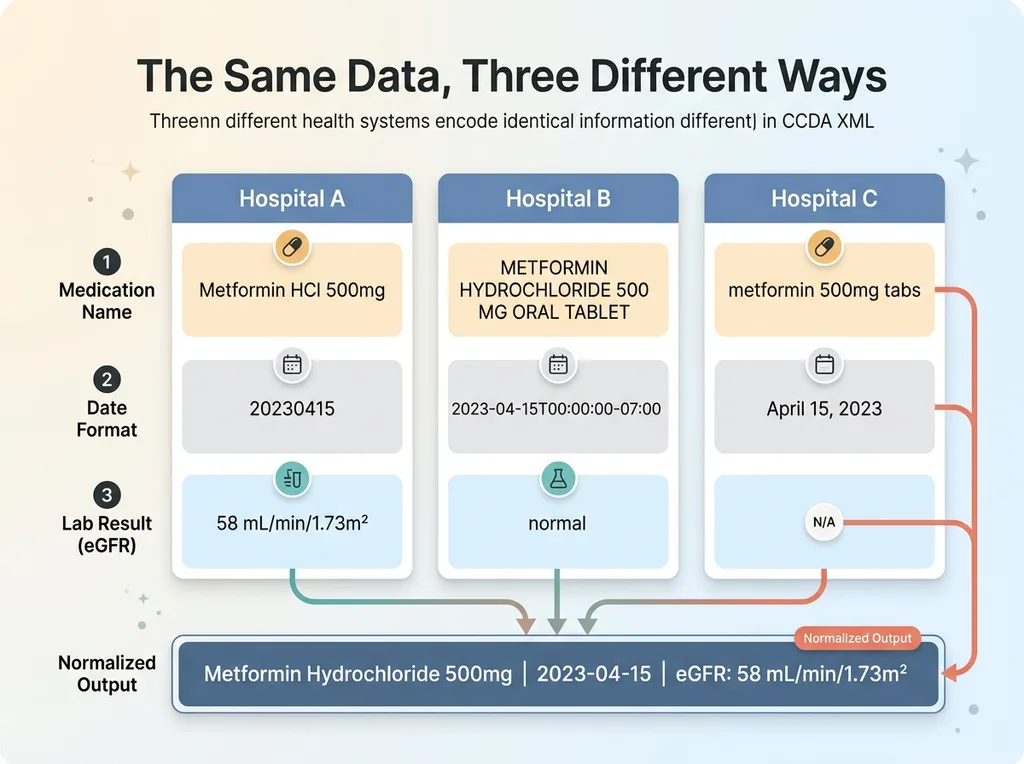
Here's what I mean by chaotic. One health system encodes dates as 20230415. Another uses 2023-04-15T00:00:00-07:00. A medication appears as "Metformin HCl 500mg" in the active medications section and as "METFORMIN HYDROCHLORIDE 500 MG ORAL TABLET" in the discharge summary. Lab results from one facility report eGFR in mL/min/1.73m² with a precise value. Another facility's CCDA just says "normal" with no numeric value at all.
Every health system implements the CCDA standard slightly differently. Some pack detailed coded entries. Others dump free-text narratives into supposedly structured sections. The "standard" is more of a suggestion.
This is why naive regex parsing breaks immediately. You can't write pattern-matching rules for a format where the patterns change depending on which hospital generated the file. You need a health record AI approach that understands clinical context — not just XML structure — to reconcile these inconsistencies across providers.
Building the CCDA Parser: Extracting Meds, Allergies, and Labs
The Extraction Pipeline
The parser is Python with lxml at its core. First pass is pure XML extraction — I walk the document tree and pull raw sections by their templateId. The medications section uses OID 2.16.840.1.113883.10.20.22.2.1.1. Allergies use 2.16.840.1.113883.10.20.22.2.6.1. Lab results, problems, procedures — each has its own identifier.
This first pass gives me raw clinical data, still messy, but isolated by category. I extract every element I can find: coded values, display names, dates, status indicators, free-text narratives, reference ranges for labs.
Normalizing the Mess
The second pass is where AI medical record parsing actually happens. Claude processes each extracted section to normalize entries across multiple source documents.
 AI Medication Normalization Process
AI Medication Normalization Process
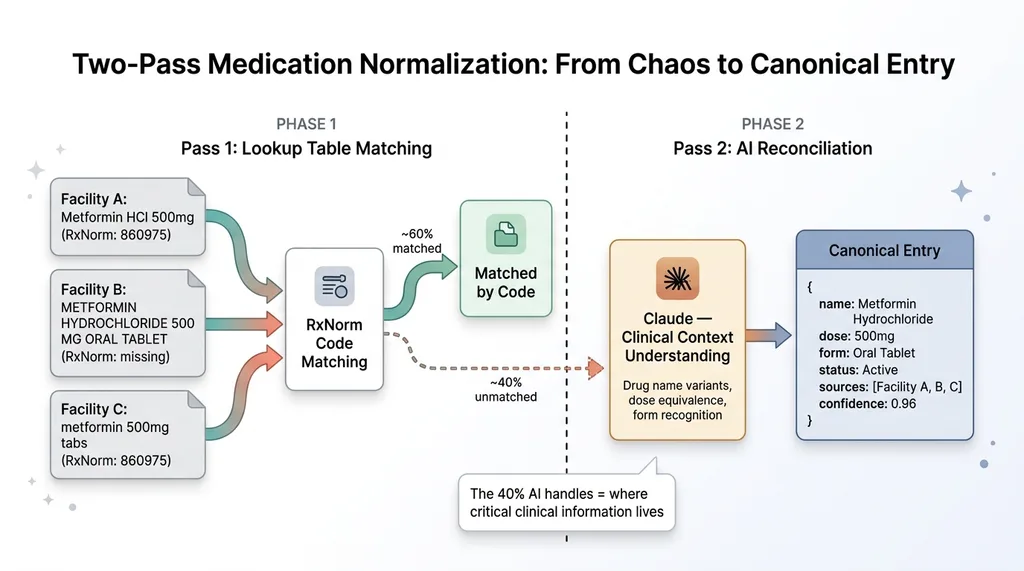
Concrete example: across three CCDA files from different facilities, the same medication appeared as:
- "Metformin HCl 500mg" (brand-style abbreviation)
- "METFORMIN HYDROCHLORIDE 500 MG ORAL TABLET" (full generic with form)
- "metformin 500mg tabs" (shorthand from a referring provider)
These are all the same medication. A lookup table approach gets you maybe 60% of the way — you can match RxNorm codes when they exist. But codes are often missing or inconsistent. Claude handles the reconciliation by understanding that these are all the same drug at the same dose, and it merges them into a single canonical entry with all source references preserved.
Same problem with labs. One facility's comprehensive metabolic panel has 14 discrete values with units and reference ranges. Another facility reports the same panel with 8 values and no reference ranges. A third facility buries the results in a narrative text block instead of coded entries.
The output is clean JSON objects — one per medication, allergy, lab result, and condition — with standardized fields: name, dose, frequency, status, date range, source document, and confidence score. This structured knowledge base is the foundation everything else builds on.
Pattern matching and lookup tables handle the straightforward cases. AI handles the 40% of edge cases that would otherwise require a human reviewer with clinical knowledge. In medical data, that 40% is where the critical information lives.
Vectorizing 30 MyChart PDFs With Voyage AI Embeddings
Why PDFs Required a Different Approach
Not everything comes as CCDA XML. Specialist visit notes, imaging reports, discharge summaries, referral letters — these are overwhelmingly PDFs. Some are generated from the EHR. Some are scanned documents. A few are practically illegible.
I had 30 PDFs from various providers. They ranged from 2-page visit summaries to 15-page discharge documents. The challenges are different from XML parsing. OCR quality varies wildly. Table layouts — especially lab panels and medication reconciliation tables — break standard text extraction. A two-column layout gets read left-to-right across both columns instead of down each one.
Chunking Clinical Documents Without Losing Context
I built an extraction pipeline that preserves document layout before chunking. Tables get extracted as structured data. Multi-column layouts get detected and handled. Headers and section markers get preserved as metadata.
 Clinical Document Chunking Strategy
Clinical Document Chunking Strategy
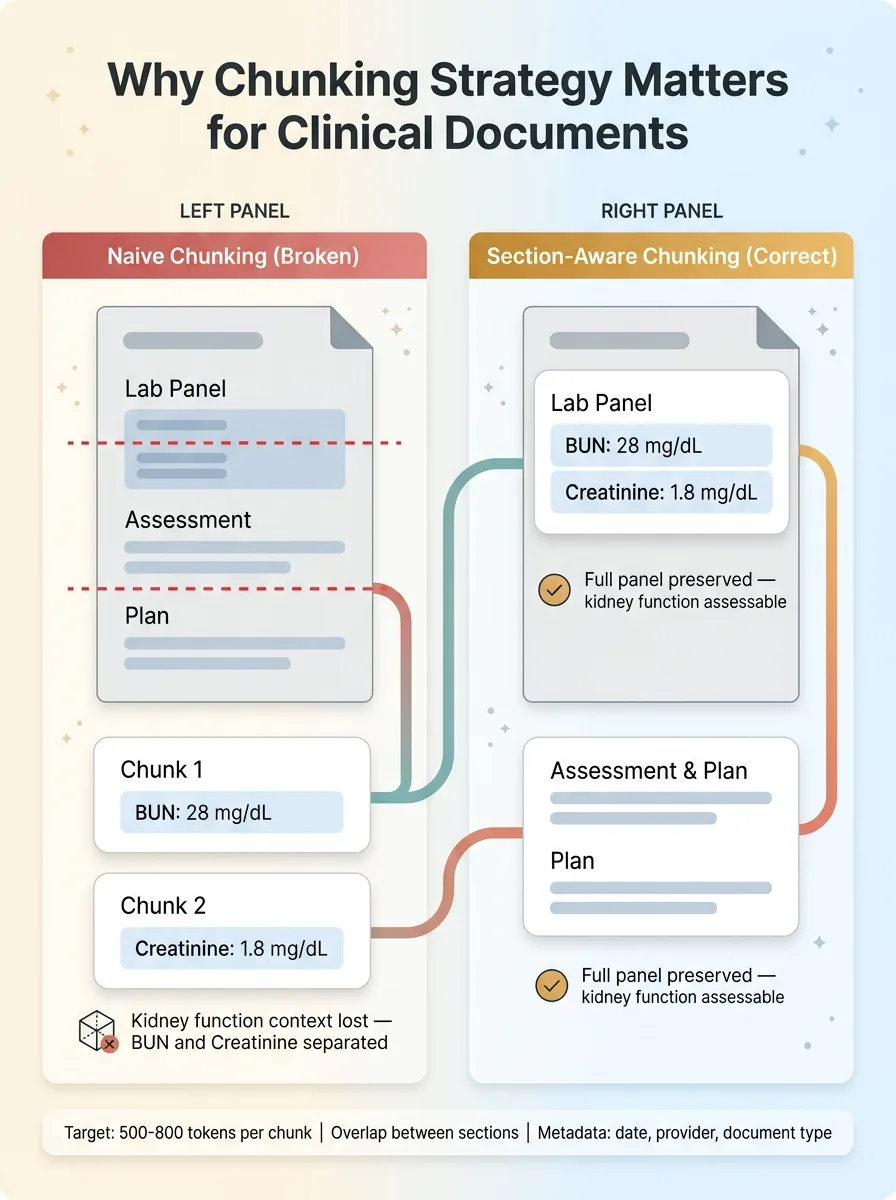
The chunking strategy matters enormously for clinical documents. You can't split a lab panel across two chunks — a BUN result in one chunk and the creatinine in another makes both useless for assessing kidney function. Assessment and plan sections need to stay together. I chunk by clinical section boundaries, with overlap, targeting roughly 500-800 tokens per chunk.
For embeddings, I tested Voyage AI against OpenAI's embedding models on this specific dataset. Voyage performed measurably better on domain-specific medical text retrieval — pulling the correct clinical context for queries about specific conditions and treatments with notably higher precision. When you're dealing with medical data extraction, embedding quality directly affects whether the system retrieves the right specialist note or gives you an unrelated visit summary.
Each chunk gets embedded and stored with metadata: document date, provider name, document type (visit note, discharge summary, imaging report), and relevant clinical context markers. From 30 documents, this produced roughly 400 chunks in the vector store. This unstructured knowledge base complements the structured CCDA data — together, they cover the full picture.
This is the same multi-model AI architecture I use across my other projects. Different models handle different parts of the pipeline because no single model excels at everything. Claude reasons over clinical content. Voyage AI handles embeddings. The architecture adapts to the task.
The RAG Pipeline: Asking Clinical Questions and Getting Real Answers
How Retrieval-Augmented Generation Works for Medical Data
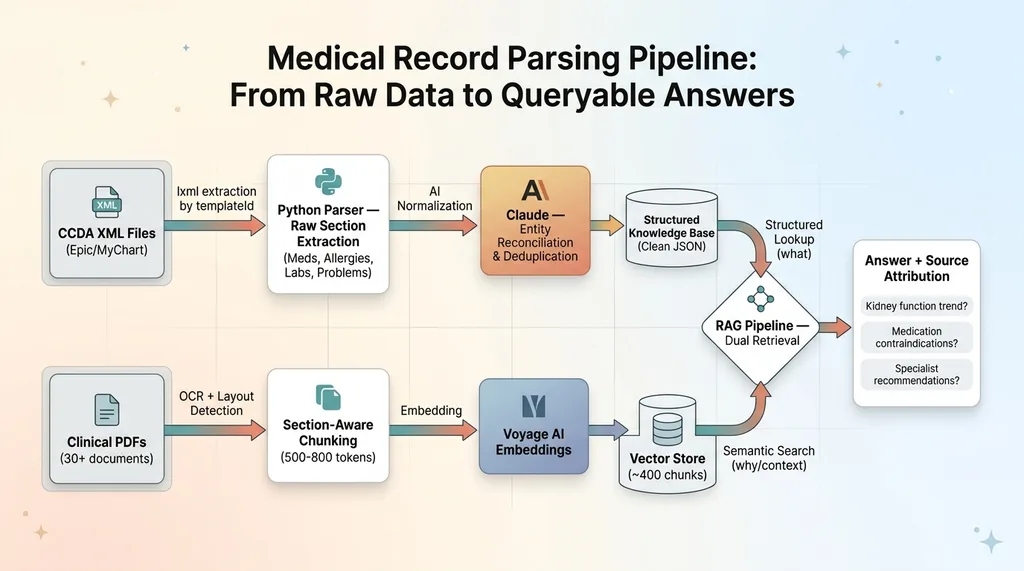
The two data sources — structured CCDA extracts and vectorized PDFs — feed into a single retrieval-augmented generation pipeline. When a query comes in, the system searches both: structured lookups for specific medications, lab values, and allergies, plus semantic search across the clinical note embeddings for context and nuance.
 End-to-End CCDA Parsing and RAG Pipeline Architecture
End-to-End CCDA Parsing and RAG Pipeline Architecture
The structured data answers "what." The unstructured data answers "why" and "what next."
What the System Can Actually Answer
Here are real queries the system handles:
"What's the kidney function trend over the last 18 months?" The system pulls eGFR and creatinine values from the structured lab data, orders them chronologically, calculates the trend, and then retrieves the nephrologist's visit notes from the vector store to provide clinical context about what the doctor said about those results.
"Are any current medications contraindicated given the allergy list?" This cross-references the normalized medication list against the allergy entries, flagging drug-class conflicts. It's not just exact matches — it catches that a documented sulfa allergy should raise a flag about certain diuretics.
"What did the cardiologist recommend at the last visit about the blood pressure medication?" Pure semantic retrieval from the vectorized visit notes, filtered by provider and date metadata.
I have to be honest about limitations. The system provides information synthesis, not medical advice. It's wrong sometimes, particularly with ambiguous clinical abbreviations — "CP" could mean chest pain, cerebral palsy, or a dozen other things depending on context. Handwritten notes that were scanned and OCR'd are unreliable.
That's why every answer includes source attribution. Every claim links back to the specific document, section, and date. A family member or caregiver can verify anything the system surfaces. This is non-negotiable for medical data — trust requires traceability.
The parsed CCDA data is also just one layer of a larger system. It feeds into what I've written about separately — a 7-specialist AI medical team architecture where different AI agents handle different aspects of health monitoring. The CCDA parser gives those agents the structured medical history they need to provide meaningful analysis.
And because this data is protected health information, the storage layer uses AES-256-GCM encryption at rest. Parsing and structuring medical data creates an obligation to protect it at a standard that exceeds the casual approach most consumer apps take. That companion piece covers the security architecture in detail.
The Healthcare Data Problem Nobody Talks About
Healthcare has a data format problem that makes every other industry I've worked in look organized. HL7v2 messages from the 1990s. FHIR APIs that are technically available but practically useless without custom integration work. CCDA exports that vary by implementation. Proprietary PDF formats. Faxed documents — in 2025. Handwritten prescription notes.
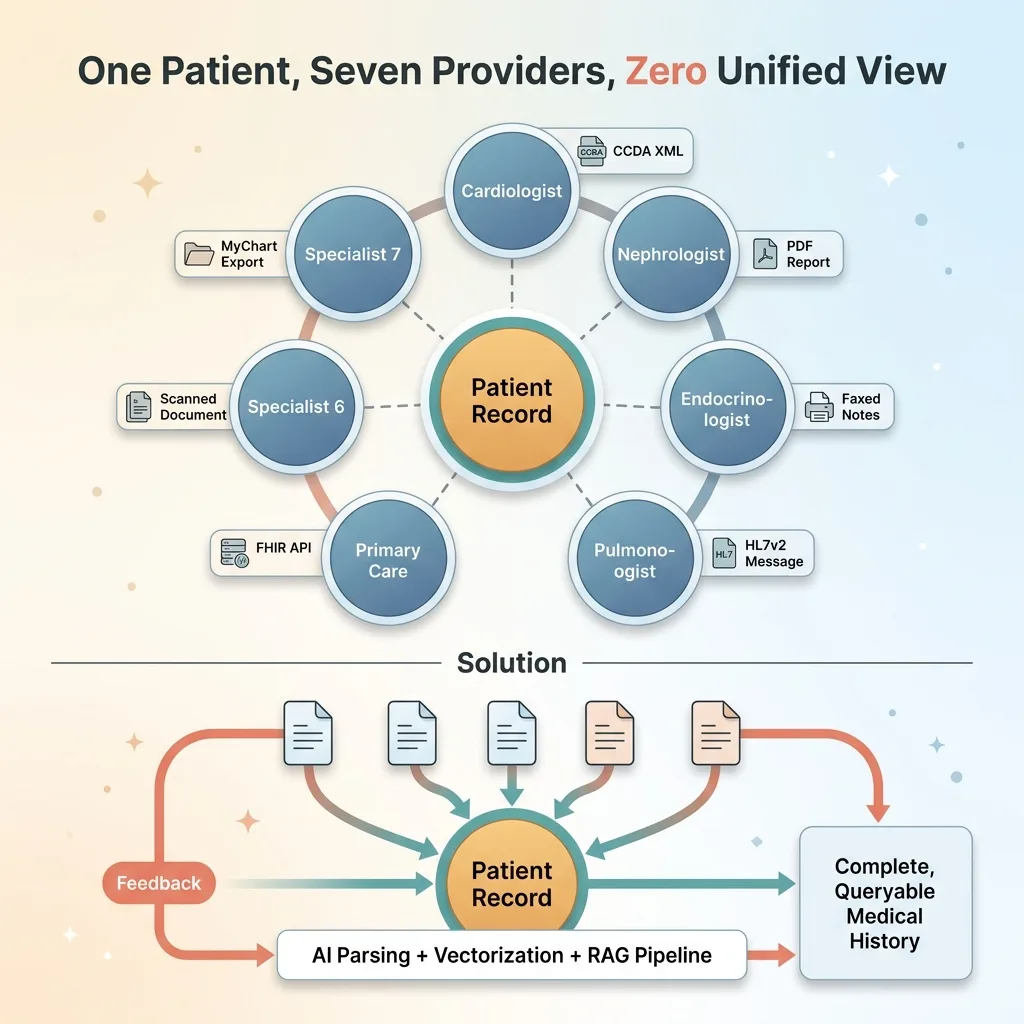 Healthcare Data Fragmentation Problem
Healthcare Data Fragmentation Problem
The promise of interoperability has been "five years away" for twenty years.
For individual patients and their families, this means critical health information is fragmented across systems that don't talk to each other. Your cardiologist doesn't know what your nephrologist prescribed last week. Your primary care doctor gets a one-paragraph referral summary instead of the full specialist workup.
AI doesn't fix the interoperability problem. But it makes it possible for a motivated technical person to build what the healthcare system won't: a unified, queryable view of one patient's complete medical history. A CCDA parser AI combined with document vectorization and RAG isn't a replacement for healthcare IT infrastructure. It's a workaround for its failure.
The pattern — messy data in, structured knowledge out — applies to any industry drowning in inconsistent document formats. Legal discovery across thousands of contracts. Financial compliance across decades of regulatory filings. Manufacturing specs from 50 suppliers in 50 formats. The data shapes change. The engineering challenge is identical.
What It Takes to Build AI Systems for Data That Fights Back
The medical records project is one example of a pattern I see everywhere: real-world data is never clean, never standard, and never ready for AI out of the box. The hard part isn't the AI model. It's the ingestion, normalization, and validation layers that make AI useful instead of dangerous.
This is what separates a weekend demo from a production system. Anyone can feed a clean dataset into Claude and get impressive-looking results. The real work is handling the 30% of records where the date format is wrong, the medication name is abbreviated, the lab units are missing, and the document structure doesn't match the standard it claims to follow.
I've built 15+ AI systems across product creation, SEO, pricing, customer service, and now healthcare. The technical foundations — Python pipelines, multi-model architectures, structured extraction, vector search — translate directly. My DTC fashion brand has 22,000+ lines of custom Python handling exactly this kind of data normalization, just for product catalogs and pricing instead of medical records.
If your business is sitting on messy, valuable data that nobody can synthesize — whether it's clinical records, financial documents, manufacturing specs, or customer data spread across six SaaS tools — that's exactly the kind of problem worth solving.
Thinking About AI for Your Business?
If any of this resonated — especially the part about valuable data trapped in formats that resist extraction — I'd like to hear about it. I do free 30-minute discovery calls where we look at your operations and identify where AI could actually move the needle for your specific situation. No slides. No pitch deck. Just an honest conversation about what's possible.
Get AI insights for business leaders
Practical AI strategy from someone who built the systems — not just studied them. No spam, no fluff.
Ready to automate your growth?
Book a free 30-minute strategy call with Hodgen.AI.
Book a Strategy Call